- Python 98%
- Shell 2%
| assets | ||
| Documentation | ||
| Example | ||
| src/renameseq | ||
| tests | ||
| .gitignore | ||
| LICENSE.txt | ||
| main.py | ||
| nuitka.sh | ||
| pyproject.toml | ||
| README.md | ||
| renameSeq.spec | ||
renameSeq
Table of Contents
Introduction
Output Folder and File Names
After completing an analisys, ABI 3500 Sequencer will generate a folder containing a max of 192 files (96 files pairs) per plate.
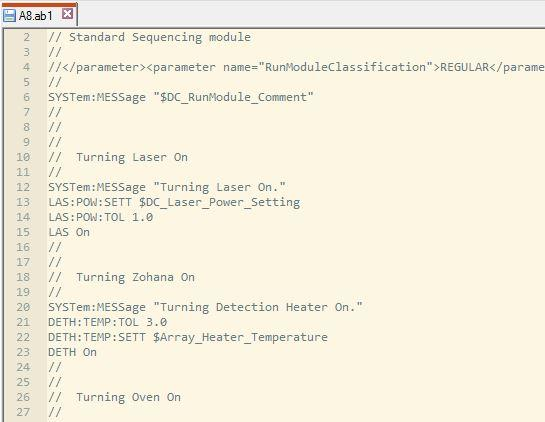
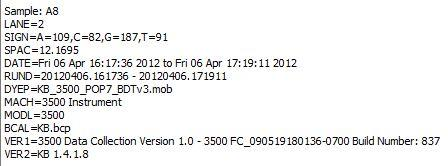
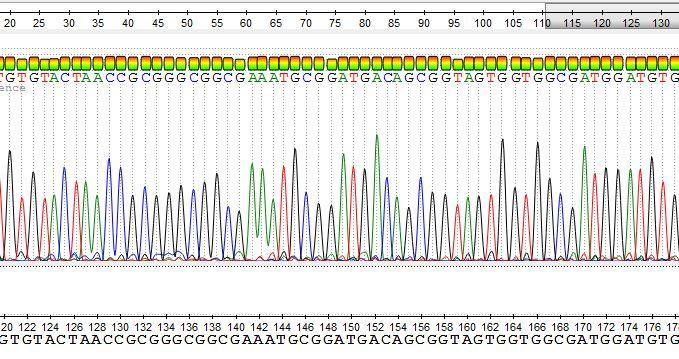
- <plate_well>.ab1 file contains a detailed report about run conditions and variations, as well as chromatographic informations and bases calls.
 |
 |
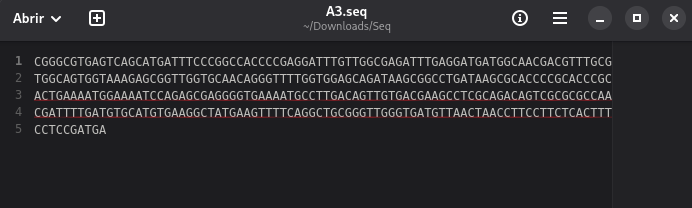
- <plate_well>.seq a plain text file containing the bases sequence read by the equipament, regardless of quality.
Note
File names will star at A1 and end at H12 for a full plate.
We will work with the Applied Biosystems Inc. Format (ab1) file. Since renameSeq will save a fasta file with the sequence found in the ab1 files. If you wish, you can compare the fasta files generated by renameSeq and those .seq files created after the analysis to make sure the final data are the same.
Traceability
Are you sure you know which sequence belong to which sample and primer used?

Good! Now...
Is it pratical? You quickly know which file has which information? Is this map of yours always in hands?
Most important of all. Do others have access to all that information, e.g. your advisor? Will it persist through time?
The most reliable way to keep track of such information is to change the original file names to something meaninful! What about the sample identification and primer used in that plate well?
The Map
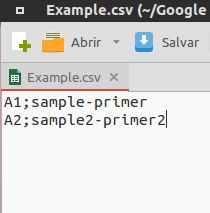 Create the file on the spreadsheet program of you preference. Name the map file as you wish - but remember - the simpler the better. 2 columns, no headers, no repeated names in any column.
Create the file on the spreadsheet program of you preference. Name the map file as you wish - but remember - the simpler the better. 2 columns, no headers, no repeated names in any column.
Save as Comma Separated Value (.csv or .txt) with semicolon as delimiter.
Fill only as many sample as you have, no need to create all the 96 if you don’t have it all. Delete the blank ones!
Caution
No special character in anywhere! Nor in the file name nor in the file content! E.g.: * , ; < > ' ^ ~ } { ) ( ] [ % $ # @ ! ? Avoid accentuation and spaces. Underscores are an excelent alternative to replace spaces if needed. You can always appeal to snake_case or CamelCase.
We are set! Let's get to the point of all this -> renameSeq application.
Installation
In a virtual environment execute:
pip install renameSeq
You may also install it using pipx or download a pre-compiled binary from the releases page and use it as a standalone executable.
Usage
Caution
ALWAYS HAVE BACKUP of the files you will submit to
renameSeqexecution. Do not expose your original single-copy files to it.renameSeqhas protections against critical mistakes, but never risk your original files. You might lose important data if so.
renameSeq is a command line interface (CLI) application. An application executed at a terminal emulator, which means black screen with a blinking cursor. It may be installed using Python's pip or used as a standalone executable.
Even though the app name has the word "rename" on it. This reflects more its intent then the way it works. For safety reasons renameSeq create new copies of the original files.
First let's stablish some good practices so the application can run smoothly:
- Do not use special characters anywhere, nor in the filenames nor in the map data.
- The new names you chose to put in your map have to be unique, no repetitions. If there are repetition renamed files will be overwritten and will not match latter, rendering them useless.
- Make sure you have the necessary permissions to execute the application (more of a Linux users concern).
- Have the sequence files (.ab1) that you want renamed in the same folder of your plate map (.csv).
- Avoid having any other files or directories inside the folder you will use to put your .ab1 files and map.
- It is not mandatory, but you should add the app to your PATH. There are diferent ways of doing it for Windows and Linux (where aliases can also be used). In case you prefere not to, pay attention to absolute and relative paths. Having the application in yout PATH allows you to use it anywhere (that you haver permissions to) in your system, by just calling it directly.
Note
In the execution examples below the app is in not in my path and I am using relative paths in a Linux system (Debian Trixie).
Once you have all set just call renameSeq fallowed by the path to your plate map file (.csv). If everything is correct a 'renamed' folder will be created inside the directory of your map with the .ab1 files renamed according to the map new names and fasta files with the sequences following the same rule.
Errors may occur and renameSeq will try its best to let you know what happend so you can correct it and restart.
After the "renaming" process renameSeq will do a checksum (SHA3-512) verification of the renamed files, if anything do not match you will be warned and should discard the renamed files, as they are not trustworthy.
License
renameseq is distributed under the terms of the LGPL-3.0-or-later license.
Additional Information
Icon downloaded from Flaticon.
Linux binary was compiled using Nuitka Standard.
Windows binary was compiled using PyInstaller.
Disclaimer
The developer of renameSeq has no affiliation of any kind with neither Applied Biosystems Inc. or Thermo Fisher Scientific Inc. Therefore renameSeq is NOT AN OFFICIAL PRODUCT from none of the companies mentioned.
Warning
This program comes with ABSOLUTELY NO WARRANTY. Use it at your own risk.


